Lectin Domain Conserved Carbohydrate-binding Site Analysis
Analysis of conserved carbohydrate-binding residues in plant lectin domains using multiple sequence alignment and WebLogo conservation visualization.
Keywords
carbohydrate binding, lectin domains, WebLogo, sequence conservation, ClustalOmega, MAFFT, plant lectins
Carbohydrate-binding Site Analysis
Objective of this section
Determine which identified lectin domains retain functional carbohydrate-binding activity by analyzing conservation of critical binding residues compared to model sequences and well-studied lectins from the literature.
Methodology overview
- Lectin domain sequences are extracted from Pfam (or relevant annotation) domain coordinates
- Untrimmed sequence alignments of lectin families with model sequences
- WebLogo3 generation for conservation visualization
- Visual inspection of conservation patterns at critical positions against the model sequence and known residues important for carbohydrate binding activity inferred from literature
Tools and databases
| Tool | Tested Version | Platform (OS) | Purpose |
|---|---|---|---|
| ClustalOmega1 | 1.2.4 | Windows/Linux/Web | Multiple sequence alignment (MSA) |
| MAFFT2 | 7.526 | Windows, Linux, macOS and Web | MSA |
| WebLogo33 | 3.7.12 | Web tool and CLI (Linux) | Conservation visualization |
Table 1. Essential software tools and databases for that can be used in this analysis. {#tbl-tools}
Key insights
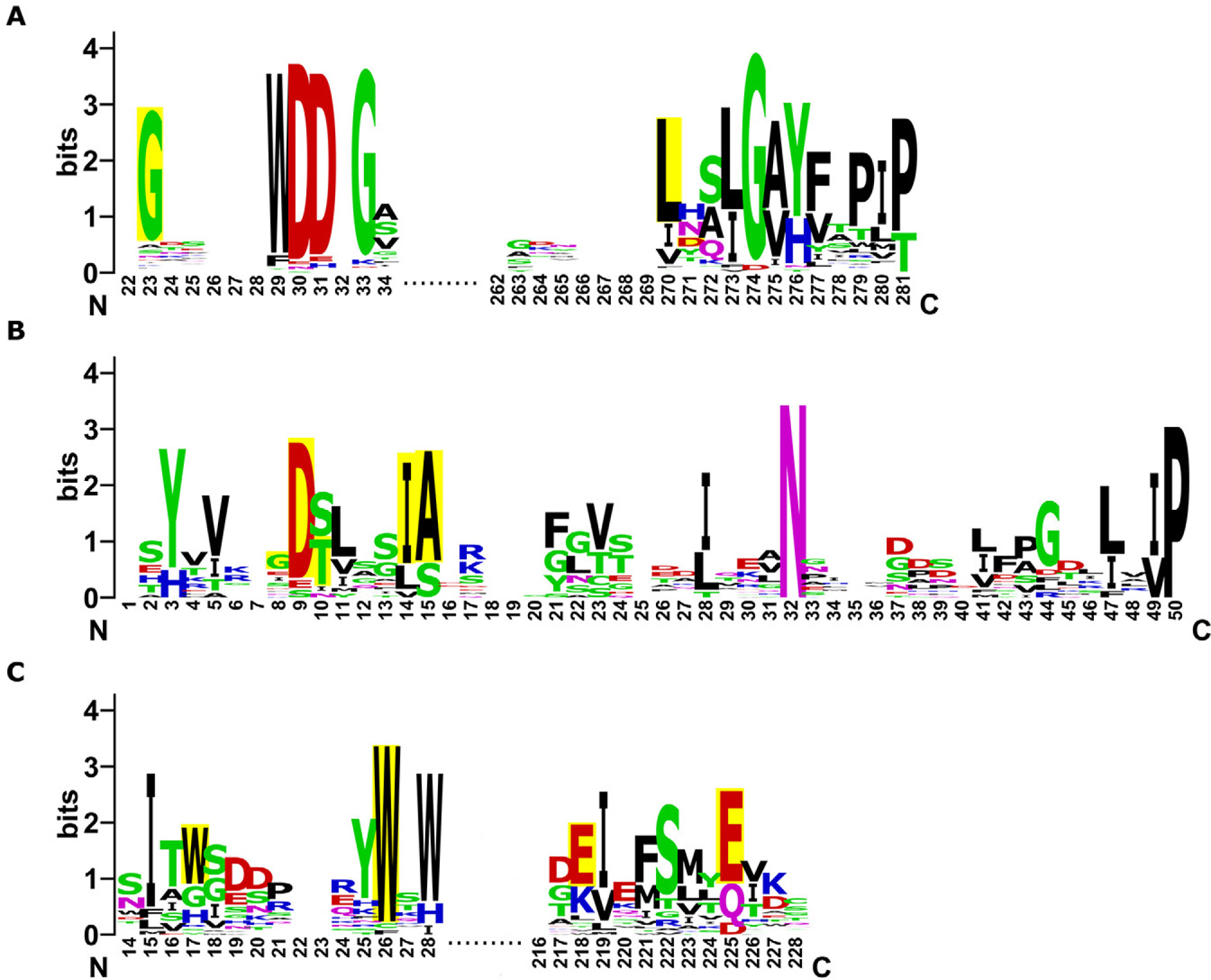
- Conservation plots showing amino acid frequency at each position with conserved residues important for carbohydrate binding activity highlighted. Each column represents a stack of letters where letter height indicates frequency and overall stack height shows sequence conservation.
- High conservation (>80%) at functional residues indicates likely activity. Missing critical residues suggests loss of activity and sequence divergence.
Key Limitations
- Alignments often show extensive gaps which makes data interpretation difficult
- Residues surrounding the binding pocket and overall protein structure can also influence specificity and activity
- Different binding sites within the same protein can show different conservation levels making functional predictions inconsistent
Key Insights
- Low sequence conservation across domains questions their functionality
- Multiple amino acid residues in binding sites can be conserved but the domain may still be non-functional
- Chimeric lectins may lose their carbohydrate-binding activity entirely
Published Studies
Phaseolus Species5, Arabidopsis thaliana4, and cucumber (Cucumis sativus)6
References
1.
Sievers, F. & Higgins, D. G. Clustal omega. Current Protocols in Bioinformatics 48, (2014).
2.
Katoh, K., Rozewicki, J. & Yamada, K. D. MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Briefings in Bioinformatics 20, 1160–1166 (2017).
3.
Schneider, T. D. & Stephens, R. M. Sequence logos: A new way to display consensus sequences. Nucleic Acids Research 18, 6097–6100 (1990).
4.
Eggermont, L., Verstraeten, B. & Van Damme, E. J. M. Genome-wide screening for lectin motifs in arabidopsis thaliana. The Plant Genome 10, plantgenome2017.02.0010 (2017).
5.
Osman, M. E. M. et al. Lectin gene families in three phaseolus species: Genome-wide identification, evolutionary analysis, pleiotropic effect, and regulation under multiple stress conditions. Journal of Molecular Evolution 94, 28–51 (2025).
6.
Dang, L. & Van Damme, E. J. M. Genome-wide identification and domain organization of lectin domains in cucumber. Plant Physiology and Biochemistry 108, 165–176 (2016).